DIMENSION REDUCTION: PRINCIPAL COMPONENTS AND
PARTIAL LEAST SQUARES
1. Principal Components Regression
> attach(Auto); library(pls)
> pcr.fit = pcr( mpg ~ . – name – origin
+ as.factor(origin), data=Auto ) #
Using all variables except name
> summary(pcr.fit)
Number of components
considered: 8
TRAINING: % variance explained
1 comps 2 comps
3 comps 4 comps 5 comps
6 comps 7 comps 8 comps
X 99.76 99.96
100.00 100.00 100.00
100.00 100.00 100.00
mpg 69.35 70.09
70.75 80.79 80.88
80.91 80.93 82.42
# The “X” row shows % of X variation contained
in the given number of PCs.
# The “mpg” row shows R2 (% of Y
variation explained) from the PC regression. The usual linear regression on all
8 variables has the same R2 as PCR that uses all 8 principal
components.
> reg = lm( mpg ~
.-name-origin+as.factor(origin), data=Auto )
> summary(reg)
Multiple R-squared: 0.8242
1a. Principal components.
# Let’s investigate the principal components,
and how much variance they explain.
> X = model.matrix( mpg ~
.-name-origin+as.factor(origin), data=Auto )
> pc = princomp(X)
> summary(pc)
Importance of components:
Comp.1 Comp.2 Comp.3 Comp.4 Comp.5
Standard deviation
854.5664182 38.860050688 1.614144e+01 3.309297e+00 1.694518e+00
Proportion of Variance
0.9975617 0.002062789
3.559039e-04 1.495959e-05 3.922297e-06
Cumulative Proportion
0.9975617 0.999624521
9.999804e-01 9.999954e-01 9.999993e-01
Comp.6 Comp.7 Comp.8 Comp.9
Standard deviation
5.242357e-01 4.162175e-01 2.443204e-01 1.110223e-16
Proportion of Variance 3.754062e-07 2.366403e-07 8.153944e-08
1.683715e-38
Cumulative Proportion
9.999997e-01 9.999999e-01 1.000000e+00 1.000000e+00
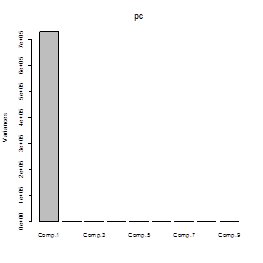
# So, Z1, the first PC, contains
99.76% of the total variation of X variables. The first two PCs together
contain 99.96%. Here is a plot of these percents called a screeplot.
> screeplot(pc)
# The actual coefficients can be obtained by
prcomp().
> prcomp(X)
PC1 PC2 PC3 PC4 PC5
(Intercept)
0.0000000000 0.0000000000 0.000000000
0.000000000 0.000000e+00
cylinders -0.0017926225 0.0133245279 -0.007294275 0.001414710
1.719368e-02
displacement
-0.1143412856 0.9457785881
-0.303312504 -0.009143349 -1.059355e-02
horsepower
-0.0389670412 0.2982553337 0.948761071 -0.043076559 -8.646402e-02
weight -0.9926735354
-0.1207516411 -0.002454212
0.001480458 3.152970e-03
acceleration
0.0013528348 -0.0348264293 -0.077006895
0.059516278 -9.944974e-01
year
0.0013368415 -0.0238516081 -0.042819254 -0.996935229 -5.549653e-02
as.factor(origin)2
0.0001308250 -0.0024889942
0.002857670 0.022100094
-9.052576e-05
as.factor(origin)3
0.0002103564 -0.0003765828
0.004796684 -0.012089823 -1.150938e-03
PC6 PC7 PC8 PC9
(Intercept) 0.0000000000 0.0000000000
0.000000e+00 1
cylinders
0.9911554803 0.1211162208
-4.909265e-02 0
displacement
-0.0146594359 -0.0006512752
4.394368e-03 0
horsepower
0.0038232742 0.0034425206
-4.435100e-03 0
weight -0.0002093216
-0.0003053766 5.729471e-06 0
acceleration
0.0168319859 0.0012233398
-1.799780e-03 0
year
-0.0001647840 0.0240346554 7.643176e-03
0
as.factor(origin)2 -0.0483462982
0.6888706846 7.229226e-01 0
as.factor(origin)3 0.1214929883
-0.7142804151 6.891098e-01 0
1b. Standardized scale
So, we see that the 1st principal component contains
a huge portion of the total variation of X variables, and it is dominated by variable
“weight”. Of course! Looking at the
data, we see that weight simply has the largest values.
> head(Auto)
mpg cylinders displacement horsepower weight
acceleration year origin
1 18
8 307 130
3504 12.0 70
1
2 15
8 350 165
3693 11.5 70
1
3 18
8 318 150
3436 11.0 70
1
4 16
8 304 150
3433 12.0 70
1
5 17
8 302 140 3449
10.5 70 1
6 15
8 429 198
4341 10.0 70
1
# For this reason, usually, X variables are
standardized first (subtract each X-variable’s mean, divide by st. deviation).
> pcr.fit = pcr( mpg ~
.-name-origin+as.factor(origin), data=Auto, scale=TRUE )
> summary(pcr.fit)
TRAINING: % variance explained
1 comps 2 comps
3 comps 4 comps 5 comps
6 comps 7 comps 8 comps
X 56.9 73.02
84.29 92.38 97.29
98.86 99.59 100.00
mpg 71.8 73.64
73.96 79.25
79.25 80.22 81.55
82.42
1c. Cross-validation
# Cross-validation. By default, this is a K-fold
cross-validation with K=10.
> pcr.fit = pcr( mpg ~
.-name-origin+as.factor(origin), data=Auto, scale=TRUE, validation="CV" )
> summary(pcr.fit)
VALIDATION: RMSEP
Cross-validated using 10 random segments.
(Intercept) 1 comps
2 comps 3 comps 4 comps
5 comps 6 comps 7 comps
8 comps
CV 7.815 4.162
4.036 4.028 3.611
3.616 3.537 3.427
3.350
adjCV 7.815 4.161
4.034 4.026 3.607
3.613 3.533 3.422
3.346
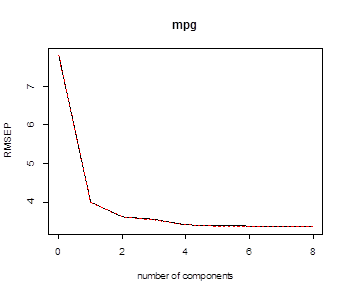
# The predicted error (by cross-validation) is
minimized by using all M=8 principal components.
# We can see the graph of root mean-squared error of prediction (or specify val.type)
> validationplot(pcr.fit)
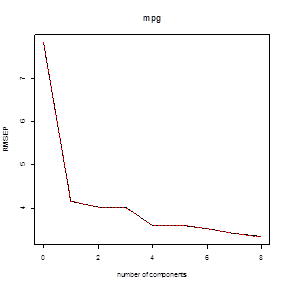
2. Partial Least Squares.
# Similar commands, just replace “pcr” with
“plsr”. M=6 components gives the lowest prediction MSE.
> pls = plsr( mpg ~
.-name-origin+as.factor(origin), data=Auto, scale=TRUE,
validation="CV" )
> summary(pls)
Cross-validated using 10 random segments.
(Intercept) 1 comps
2 comps 3 comps 4 comps
5 comps 6 comps 7 comps
8 comps
CV 7.815 3.994
3.616 3.540 3.395
3.379 3.351 3.364
3.362
adjCV 7.815 3.992
3.612 3.535 3.390
3.376 3.345 3.359
3.357
TRAINING: % variance explained
1 comps 2 comps
3 comps 4 comps 5 comps
6 comps 7 comps 8 comps
X 56.73 68.84
80.75 84.08 93.48
94.88 99.33 100.00
mpg 74.32 79.37
80.29 81.71 82.00
82.35 82.38 82.42
> validationplot(pls)